Discovery Tools
From clustering and diversity analysis for chemical libraries to 2D and 3D molecular screening
JKlustor
JKlustor Summary
JKlustor - Clustering and diversity analysis for chemical libraries
JKlustor is a suite packed with clustering methods, that performs similarity and structure based clustering of compound libraries and focused sets - in both hierarchical and non-hierarchical fashions. The suite also carries out diversity calculations and library comparisons based on molecular fingerprints and other descriptors. It is an essential tool in combinatorial chemistry, virtual library design, and other areas where large numbers of compounds are analyzed.

Features of JKlustor
Similarity based clustering
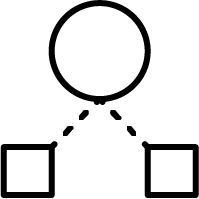
Hierarchical method:
Ward’s minimum variance method, speeded up with Murtagh’s reciprocal nearest neighbour algorithm, creates tight and well separated clusters. It's recommended to use it with smaller data sets, like focused libraries with less than 100,000 structures. More on Ward clustering
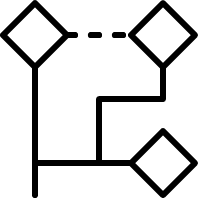
Non-hierarchical method:
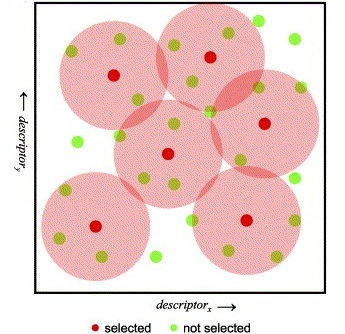
The Sphere Exclusion clustering is based on fingerprints and/or other numerical data, it can easily cope with millions of structures and it is suitable for diverse subset selection. K-means cluster analysis method aims to find the center of natural clusters in the input data in a way that minimizes the variance within each cluster. Finally, the Jarvis-Patrick (Jarp) method uses a nearest neighbor approach and performs variable-length clustering of chemical databases with hundreds of thousands of structures contained.
Features of JKlustor
Structure based clustering

Hierarchical methods:
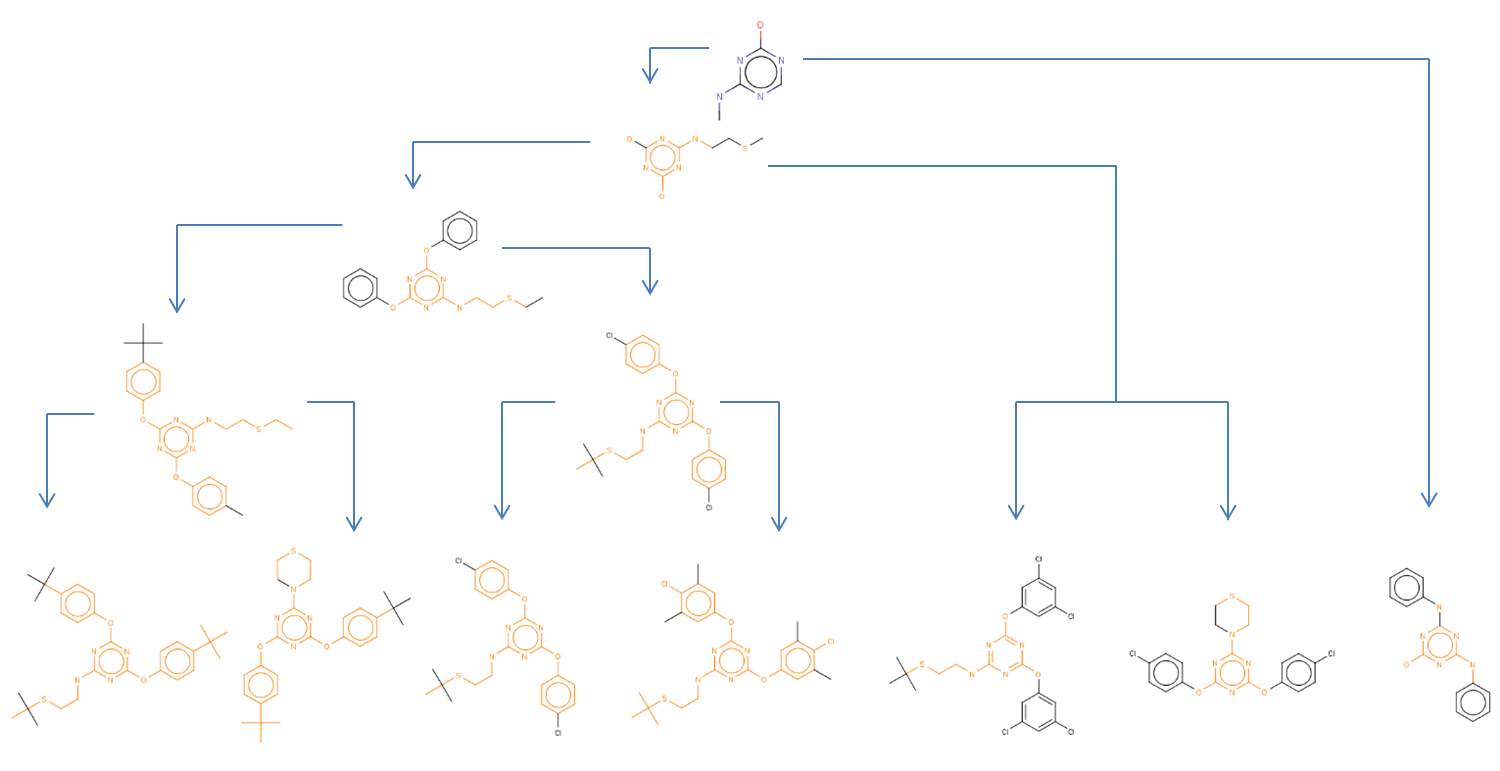
LibraryMCS identifies the largest substructure shared by several molecular structures. It uses the hierarchical representation of clusters (dendograms), and it vizualizes an alternative tree and table view too. MCS profiling helps scientists explore screening results to quickly identify novel scaffolds and new examples of active compound families. The hierarchical SAR table enables viewing of clusters and associated non-structural data. R-group decomposition can be also performed using the MCS as the core structure for each cluster.
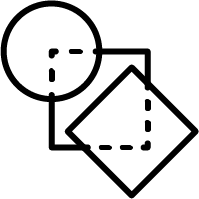
Non-hierarchical method:
JKlustor makes clustering available for pre-generated use Bemis-Murcko frameworks structures, and therefore provides a convenient and quick way towards analyzing large databases with millions of compounds.
Features of JKlustor
Extending capabilities with descriptors and command line tools
JKlustor can use Chemaxon’s proprietary chemical and pharmacophore fingerprint technology, and also other user defined descriptors such as BCUT. For example, predicted or measured phys-chem properties (like pKa, logP etc.) can also help with clustering.
Command line tools in JKlustor:
- Diversity analysis - with Compr command line it generates different types of similarity or dissimilarity comparisons within a dataset (also see Diversity Set Selection) .
- GenerateMD - generates molecular descriptors for molecular stuctures.
- Jarp - clustering by a modified Jarvis-Patrick method.
- Ward - clustering by Ward's hierarchic clustering method using the RNN approach.
- LibMCS - Maximum Common Substructure based hierarchical clustering.
- CreateView - composing a new SDfile from an SDfile and a data table - It's useful for viewing the clustered results.

JKlustor Benefits
Availability
JKlustor tools can be called upon from the command line or from the API of JChem. JKlustor runs on many operating systems and can integrate with many database engines. Full Java and .NET integration is supported, as well as, connected to Oracle, MySQL, MS SQL Server, DB2, PostgreSQL, Access, etc. databases. The LibMCS element comes with a standalone GUI that allows users to browse/navigate through large sets of data. Furthermore, maximum common edge sub-graph (MCES) and maximum common substructure (MCS) clustering methods are also available as Chemaxon components, for both, KNIME and Pipeline Pilot workflow management systems.
Screen Suite
Screen Suite Summary
Screen Suite - Toolkit for 2D and 3D molecular screening
The Screen Suite is a ligand-based high throughput virtual screening package that provides powerful tools for chemical (2D) and shape (3D) similarity searches of large molecular libraries. The suite provides several different sets of fingerprints, such as the Chemaxon chemical fingerprint, pharmacophore fingerprint, ECFP/FCFP, dissimilarity metrics and their optimization to tune your processes for optimal search results. Commercially the suite is available as 2D Descriptor package, Screen2D (including our chemical fingerprint) and Screen3D. It is also available as a command line tool, from Java and .NET API and workflow tools like KNIME or Pipeline Pilot. Screen Suite is a progenitor of the novel MadFast similarity search engine, where descriptor generation and similarity search using multiple metrics are also available.

Screen Suite Features
Flexible descriptor accessibility in Screen Suite
The Screen2D Descriptor package contains fingerprint generators for chemical structures, pharmacophores, extended connectivity (ECFP) and functional class (FCFP) descriptors. It can be used to generate and tune fingerprints for use in any screening process. Descriptor generation is supported through Chemaxon’s Chemical Terms scripting language, to create complex fingerprints which consider single or combined molecular attributes that can be based on topological or physicochemical properties like partition coefficients, hydrogen bonding donor-acceptor accessibility or acidic properties.
Screen Suite Features
Diverse fingerprints
Chemaxon's screening technology heavily relies on the available chemical fingerprints and molecular descriptors:
- BCUT descriptors - used in diversity analyses and QSAR; based on the connectivity table of molecules with physchem properties like atomic charge, HBDA and polarizability
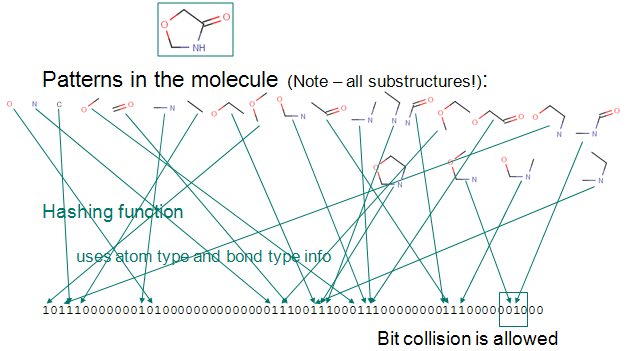
- Chemical fingerprint - used for structural searches in databases, similarity search or diversity analysis of compound libraries; it is a path-based bit-string descriptor describing molecular structures
- Pharmacophore fingerprints - atom-based fingerprints characterizing binding related structural or physchem properties that are thought to be responsible for pharmacophore activity
- Circular fingerprints - in contrast to path-based fingerprints ECFP/FCFP are not substructure preserving descriptions, but are highly relevant for full structure and similarity searches.
Screen Suite Features
Screen 2D
Screen 2D is a robust platform to perform chemical similarity searches on large molecular databases. It offers a fast process with up to 100,000 compounds per second being screened on a single desktop PC. Modularity of the tool allows to use own descriptor sets. Screen 2D comes with a set of metrics to measure dissimilarity between compounds:
- Euclidean
- Tanimoto
- Tversky
- Dice
Screen 2D supports metrics optimization by selecting appropriate training sets. The optimization step increases enrichment significantly, giving smaller, more focused hit sets. It is available through API and command line, via Pipeline Pilot and KNIME connectors, and some features are implemented within Instant JChem and JChem for Office desktop applications.
Screen Suite Features
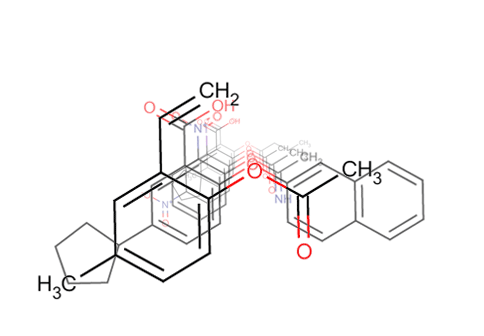
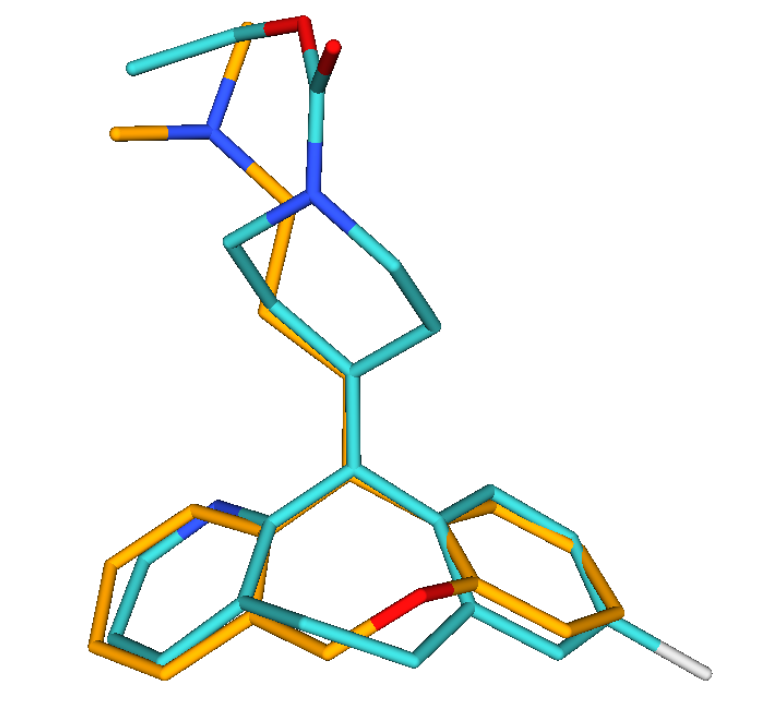
Screen 3D
Screen Suite includes 3D shape similarity based searching in addition to topology based screening. It introduces a novel flexible alignment technique, which provides reasonable alternative hit results as compared to previous 3D screen engines. The key feature of Screen 3D is definitely its high speed, however, it also compares very well in providing better enrichment in the first 1% of the hits than most alternative software. Scalability of the tool makes it suitable to search in very large molecular databases.
Screen 3D can be used to carry out different shape similarity searches. Beyond fully flexible matching it can align flexible database compounds to rigid query molecules (rigid-flexible). Alternatively it is capable of aligning rigid database compounds to rigid query molecules (rigid-rigid) using on-the-fly generated conformation ensembles. Benefits of Screen3D can also be exploited as a plugin in Design Hub idea assessment and management platform.
Knowledge Hub
Resources
Learn more about Discovery Tools
Documentation
API Docs

Related Products
Marvin
Full featured chemical editor for all platforms
Chemical Naming
Convert chemical names into structures
Markush Technology
Smart assistant for patent claim drafting and Markush analysis
Chemical Structure Representation
Standardization and correction of chemical structures
Chemicalize
Calculate properties instantly, search chemical data, and draw molecules online
Compound Registration
Normalize, check, validate and register chemical compounds
Reactor
High performance virtual synthesis engine
JChem for Office
Chemical structure handling, data analysis, visualization and reporting capabilities within MS Office
Design Hub
A single platform that connects scientific rationale, compound design and computational resources
JChem Engines
Search through tens of millions of chemical compounds and receive relevant query hits in seconds.
Calculators and Predictors
Execute high quality physico-chemical calculations and predictions.
Compliance Checker
Identify controlled substances with Compliance Checker and assign HS tariff codes with cHemTS - the easy way to comply with chemical regulations.
Discovery Tools
From clustering and diversity analysis for chemical libraries to 2D and 3D molecular screening
Instant JChem
Create, explore and share chemical data
ChemCurator
Computer-assisted chemical information extraction and analysis

